
Primate and human brain evolution
With humans as the outlier, primates are an excellent model system for investigating the link between brain evolution and the evolution of cognition.
Selected publications:
Smaers JB & Vanier DR (2019) Brain size expansion in primates and humans is explained by a selective modular expansion of the cortico-cerebellar system. Cortex, 118: 154-164.
Smaers JB, Gómez-Robles A, Parks AN & Sherwood CC (2017) Exceptional evolutionary expansion of the prefrontal cortex in great apes and humans. Current Biology. 27: 714-720.
Illustration by Javier Lazaro (http://www.lazaroillustration.com/)
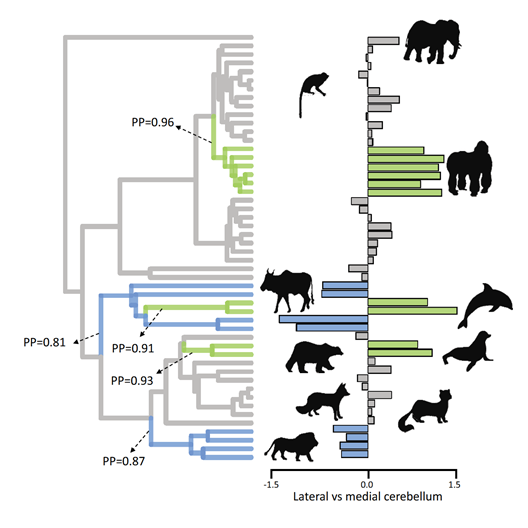
Mammalian brain evolution
Several mammalian clades have independently evolved higher cognition. Such instances of parallel and convergent evolution provide exceptional opportunities to disentangle the patterns and processes that shape brain evolution.
Selected publications:
Smaers JB, Rothman RS, et al (2021) The evolution of mammalian brain size. Science Advances. 7: eabe2101.
Smaers JB, Turner AH, Gómez-Robles A, & Sherwood CC (2017) A cerebellar substrate for cognition evolved multiple times independently in mammals. eLife, 7: e35696.

Vertebrate brain evolution
Because different classes of vertebrates display widely varying neural architecture, comparisons within and across different classes provide fundamental information on the basic principles of brain evolution.
Selected publications:
Ksepka DT, Balanoff AM, Smith NA, … Jarvis ED & Smaers JB. Tempo and pattern of avian brain evolution. Current Biology. 30: 2026-2036.
Balanoff A, Smaers JB & Turner A. Modular evolution and the influence of flight on neuroanatomical variation within theropod dinosaurs. Journal of Anatomy. 229: 204-214.